(1-1)
 |
 |
 |
Z-Score:
25.6
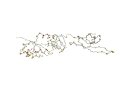
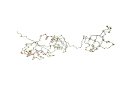
AR: 200 RMSD: 2.8A SI: 90% |
|
| 1m1g A (len 240) | 1m1g
B (len 242) |
DaliLite result | ||
| site 1m1A LEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES code --0000000000QBQAAAAAAAAAAAAABRRQABGR0000RG00000000RRG0R0000RG0000000000RR00000000RRG000R00 LCS 0000000000Q QAAAAAAAAAAAAABRRQABGR000 RG00000000 RG0R0000RG0000000000R 00000000RRG000R00 code --R00000000000QGQAAAAAAAAAAAAABRRQABGR000ORG000000000RG0R0000RG0000000000R000000000RRG000R00 1m1B QELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES site site 1m1A VEGD TCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIK R GV code 00R0 QAABG0000G0RG0000QABR000000000QB00RG00000O00QAAAAAAABG0RG000000RRG00000QAAAAAAAAAAB 0 00 LCS 00R Q ABG0000 0RG0000Q BR0000000 00RG00000 00QAAAAAAABG0RG000000RRG00000QAAAAAAAA B 0 00 code 00RGRQ ABG000000RG0000QGBR0000000R3PG00RG00000000QAAAAAAABG0RG000000RRG00000QAAAAAAAAB1BG0G00 1m1B VEGDTC VNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRGVKP site site ++ ++++++++ 1m1A KPSKVEFEKGDQVRVIEGPFMNFTGTVEEVHPEKRKLTV M I SI FGRMTPVELDFD QVEKI code RO00R0000R00000R00QB0R00000O0000QBR0000 0 0 00 RRGR000000QA G00-- LCS 00 0000R00000R00QB0R0000 R0000 0 0 00 RR R000000QA G 0 code 0000000R00000R00QB0R0000 R0000RRRRG0R0O00RRR0R000000QABG 0-- 1m1B SKVEFEKGDQVRVIEGPFMNFTGT VEEVHPEKRKLTVMISIFGRMTPVELDFDQV EKI site ++++++++++ *RATIO : lcs/trimmed_seq_a = 0.881 (= 208/ 236), lcs/trimmed_seq_b = 0.874 (= 208/ 238) |
| Structural alignment by ComSubstruct + manual inspection |
| DSSP ..LLLEEEEEEELLLLHHHHHHHHHHHHHHHLLLLLEEEEELLLEEEEEEEELLLEEEEEELLLLLEEEEEELLLLEEEEEEELLEEEEEEL 1m1A ..LEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1B qeLEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES DSSP llLLLEEEEEEELLLLHHHHHHHHHLHHHHHLLHHHEEEEELLLEEEEEEEELLLEEEEEELLLLLEEEEEELLLLEEEEEELLLLEEEEEL DSSP LLLLHHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHHLLLEEEELEELLEELEELHHHHHHHHHHHHL.LL 1m1A VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKR.GV |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1B VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRgVK DSSP LLLLHHHLLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHLLLLEEEELEELLEELEELHHHHHHHHLLLLLlLL DSSP LLLL.LLLL..........................LLLEEEelllllllleeeeeeeelllleeeeeeeelleeeEEEEELlleeel 1m1A KPSK.VEFE..........................KGDQVRviegpfmnftgtveevhpekrkltvmisifgrmtPVELDFdqveki | 1m1B PSKVeFEKGdqvrviegpfmnftgtveevhpekrkLTVMIS......................ifgrmtpveldfDQVEKI...... DSSP LLLLlLLLLleeeelllllllleeeelllllllllEEEEEE......................elleeeeeeelhHHEEEL...... *Z-score 25.6 AR 200 RMSD 2.8A SI 90% |
| Structural alignment by DaliLite |
(1-2)
 |
 |
 |
Z-Score:
25.1
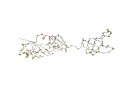
AR: 202 RMSD: 3.0A SI: 89% |
|
| 1m1g A (len 240) | 1m1g
D (len 239) |
DaliLite result | ||
| site 1m1A LEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES code --0000000000QBQAAAAAAAAAAAAABRRQABGR0000RG00000000RRG0R0000RG0000000000RR00000000RRG000R00 LCS 000000000QBQAAAAAAAAAAAAABR QABG 0000RG00000000R 0 0000RG0000000000R 00000000RRG000R00 code --000000000QBQAAAAAAAAAAAAABRPQABG00000RG00000000R00000000RG0000000000R000000000RRG000R00 1m1D EKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES site site 1m1A VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIK RGV code 00R0QAABG0000G0RG0000QABR000000000QB00RG00000O00QAAAAAAABG0RG000000RRG00000QAAAAAAAAAAB 000 LCS 00R0QAABG0000G0RG000 QABR00000000 Q 00 G00000 00QAAAAAAABG0RG000000RRG00000QAAAAAAAAAAB 000 code 00R0QAABG0000G0RG000RQABR00000000RQG000G00000000QAAAAAAABG0RG000000RRG00000QAAAAAAAAAABG000 1m1D VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRGVK site site ++++++++++ 1m1A KPSKVEFEKG DQVRVIEGPFMNFTGTVEEVHPEKRKLTVMISIFGRMTPVELDFD QVEKI code RO00R0000R 00000R00QB0R00000O0000QBR00000000RRGR000000QA G00-- LCS R 00R 00 R 00000R00QB0R00000 0000QBR00 0 00 RRGR000000QA G 0 code 1R 00R 00 R000000R00QB0R0000000000QBR00R0R00RRRGR000000QABG 0-- 1m1D PS KVE FE KGDQVRVIEGPFMNFTGTVEEVHPEKRKLTVMISIFGRMTPVELDFDQV EKI site ++++++++++ *RATIO : lcs/trimmed_seq_a = 0.915 (= 216/ 236), lcs/trimmed_seq_b = 0.919 (= 216/ 235) |
| Structural alignment by ComSubstruct + manual inspection |
| DSSP lLLEEEEEEELLLLHHHHHHHHHHHHHHHLLLLLEEEEELLLEEEEEEEELLLEEEEEELLLLLEEEEEELLLLEEEEEEELLEEEEEEL 1m1A lEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1D .EKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES DSSP .LLEEEEEEELLLLHHHHHHHHHHHHHHLLLHHHEEEEELLLEEEEEEEELLLEEEEEELLLLLLEEEEELLLLEEEEEELLLLEEEEEL DSSP LLLLHHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHHLLLEEEELEELLEELEELHHHHHHHHHHHHLLL 1m1A VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRGV |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1D VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRGV DSSP LLLLHHHLLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHHLLLEEEELEELLEELLLLHHHHHHHHHHHHHLL DSSP L.lLLLLLL...........................LLLEEEellllllllEeeeeeeelllleeeeeeeelleeeEEEEELlleeel 1m1A K.pSKVEFE...........................KGDQVRviegpfMNFTgtveevhpekrkltvmisifgrmtPVELDFdqveki | 1m1D KpsKVEFEKgdqvrviegpfmnftgtveevhpekrkLTVMIS.....iFGRM.................tpveldfDQVEKI...... DSSP LllLLLLLLlleeellllllllleeeeeeeelllleEEEEEE.....eLLEE.................eeeeeelLLLEEL...... *Z-score 25.1 AR 202 RMSD 3.0A SI 89% |
| Structural alignment by DaliLite |
(1-3)
 |
 |
 |
Z-Score:
24.0
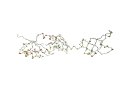
AR: 238 RMSD: 6.2A SI: 100% |
|
| 1m1g A (len 240) | 1npp
B (len 241) |
DaliLite result | ||
| site 1m1g LEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQ GKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES code --0000000000QBQAAAAAAAAAAAAABRRQABGR0000RG00000000R RG0R0000RG0000000000RR00000000RRG000R00 LCS 0000000000QBQAAAAAAAAAAAAABR QABGR0000RG0000000 R R 0 0000RG0000000000R 00000000RRG000R00 code --00000000000QBQAAAAAAAAAAAAABRHQABGR0000RG0000000 R1R0000000RG0000000000R000000000RRG000R00 1npp ELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIR AQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES site site 1m1g VEGD TCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQI KRGV code 00R0 QAABG0000G0RG0000QABR000000000QB00RG00000O00QAAAAAAABG0RG000000RRG00000QAAAAAAAAAA B000 LCS 00R QA BG0000 0RG0000QABR0000000 0Q 00RG00000000QAAAAAAABG0RG000000RRG00000QAAAAAAAAAA B code 00RGRQA BG000000RG000RQABR000000030QG00RG00000O00QAAAAAAABG0RG000000RRG00000QAAAAAAAAAAAB 1npp VEGDTCV NAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKR site site ++++++ ++++ 1m1g KPSKVEF EKGDQVRVIEGPFMNFTGTVEEVHPEKRKL TVMISIFGRM TPVELDFD QVEKI code RO00R00 00R00000R00QB0R00000O0000QBR00 000000RRGR 000000QA G00-- LCS R R00 00R00000 00QB0R00000 0 R00 000000RR 000000QA G 0 code RR R00RRRR00R00000000QB0R00000R0 R00RRRG000000RR000000000QABG 0-- 1npp GV KPSKVEFEKGDQVRVIEGPFMNFTGTVEE VHPEKRKLTVMISIFGRMTPVELDFDQV EKI site ++++++++++ *RATIO : lcs/trimmed_seq_a = 0.886 (= 209/ 236), lcs/trimmed_seq_b = 0.882 (= 209/ 237) |
| Structural alignment by ComSubstruct + manual inspection |
| DSSP .LLLEEEEEEELLLLHHHHHHHHHHHHHHHLLLLLEEEEELLLEEEEEEEELLLEEEEEELLLLLEEEEEELLLLEEEEEEELLEEEEEEL 1m1A .LEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1B eLEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES DSSP lLLLEEEEEEELLLLHHHHHHHHHHHHHHHLLHHHEEEEELLLEEEEEEEELLEEEEEEELLLLLEEEEEELLLLEEEEEEELLEEEEEEL DSSP LLLLHHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHHLLLEEEELEELLEELEELHHHHHHHHHHHHLLL 1m1A VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRGV |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1B VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRGV DSSP LLLLHHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHLLLLEEEELEELLEELLLLHHHHHHHHHHHHHLL DSSP LllLLLLLLLLEEEELLLLLLLLEEEEEEEELLLLEEEEEEEELLEEEEEEEELLLEEEL 1m1A KpsKVEFEKGDQVRVIEGPFMNFTGTVEEVHPEKRKLTVMISIFGRMTPVELDFDQVEKI | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1B KpsKVEFEKGDQVRVIEGPFMNFTGTVEEVHPEKRKLTVMISIFGRMTPVELDFDQVEKI DSSP LllLLLLLLLLEEEELLLLLLLLEEEEEELLLLLLEEEEEEEELLEEEEEEEELLLEEEL *Z-score 24.0 AR 238 RMSD 6.2A SI 100% |
| Structural alignment by DaliLite |
(1-4)
 |
 |
 |
Z-Score:
25.6
AR: 206 RMSD: 16.0A SI: 100% |
|
| 1m1g A (len 240) | 1npp
D (len 240) |
DaliLite result | ||
| site 1m1g LEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES code --0000000000QBQAAAAAAAAAAAAABRRQABGR0000RG00000000RRG0R0000RG0000000000RR00000000RRG000R00 LCS 0000000000QBQAAAAAAAAAAAAABR QABG 0000RG00000000RRG0R0000RG0000000000R 00000000RRG000R00 code --G000000000000QBQAAAAAAAAAAAAABR1QABGO0000RG00000000RRG0R0000RG0000000000R000000000RRG000R00 1npp VQELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES site site 1m1g VE GDTCVNAPPISKPGQKITCKENKTEAKIVL DNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRGV code 00 R0QAABG0000G0RG0000QABR0000000 00QB00RG00000O00QAAAAAAABG0RG000000RRG00000QAAAAAAAAAAB000 LCS 00 R QA BG0000 0RG0000QABR0000000 00 00RG00000 00QAAAAAAABG0RG000000RRG00000QAAAAAAAAA B code 00BGR QA BG000000RG0000QABR0000000R000 00RG00000R00QAAAAAAABG0RG000000RRG00000QAAAAAAAAA BRR 1npp VEGDT CV NAPPISKPGQKITCKENKTEAKIVLDNKI FPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQ IKR site site ++++++ ++++ 1m1g KPSKVEF EKG DQVRVIEGPFMNFTGTVEEVH PEKRKLTVM ISIFGRMT PVELDFDQVEKI code RO00R00 00R 00000R00QB0R00000O000 0QBR00000 000RRGR0 00000QAG00-- LCS R 00R0 00R 00000 00QB0R00000 000 0 R 0000 000RR 0 00000 00 code R 00R0R1OG00RG00000 00QB0R00000R00010 RG00003000RR 0300000 00G-- 1npp G VKPSKVEFEKGDQVRVI EGPFMNFTGTVEEVHPE KRKLTVMISIFG RMTPVEL DFDQV site +++ +++++++ *RATIO : lcs/trimmed_seq_a = 0.894 (= 211/ 236), lcs/trimmed_seq_b = 0.894 (= 211/ 236) |
| Structural alignment by ComSubstruct + manual inspection |
| DSSP ...LLLEEEEEEELLLLHHHHHHHHHHHHHHHLLLLLEEEEELLLEEEEEEEELLLEEEEEELLLLLEEEEEELLLLEEEEEEELLEEEEEEL 1m1g ...LEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1npp vqeLEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVVES DSSP lllLLLEEEEEEELLLLHHHHHHHHHHHHHHLLLHHHEEEEELLLEEEEEEEELLEEEEEEELLLLLEEEEEELLLLEEEEEEELLEEEEEEL DSSP LLLLHHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHHLLLEEEELEELLEELEELHHHHHHHHHHHHLll 1m1g VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRgv |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1npp VEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRgv DSSP LLLLLHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHLLLLEEEELEELLEELEELHHHHHHHHHLLLLll DSSP lllllllLLLLEEEElllllLLLEEEEEeeelllleeeEEEEELLEEEEeeeellleeel 1m1g kpskvefEKGDQVRViegpfMNFTGTVEevhpekrkltVMISIFGRMTPveldfdqveki |||||||| |||||||| ||||||||||| 1npp kpskvefEKGDQVRViegpfMNFTGTVEevhpekrkltVMISIFGRMTP...veldfdqv DSSP lllllllLLLLEEELlllllLLLEEEEEellllllleeEEEEELLEEEE...eeelllll *Z-score 25.6 AR 206 RMSD 16.0A SI 100% |
| Structural alignment by DaliLite |
(1-5)
 |
 |
 |
Z-Score:
27.7
AR: 230 RMSD: 16.3A SI: 98% |
|
| 1m1g B (len 242) | 1npp
D (len 240) |
DaliLite result | ||
| site 1m1g QELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVV code --R00000000000QGQAAAAAAAAAAAAABRRQABGR000ORG000000000RG0R0000RG0000000000R000000000RRG000R LCS 00000000000Q QAAAAAAAAAAAAABR QABG 000 RG00000000 RG0R0000RG0000000000R000000000RRG000R code --G000000000000QBQAAAAAAAAAAAAABR1QABGO0000RG00000000RRG0R0000RG0000000000R000000000RRG000R 1npp VQELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVV site site 1m1g ESVEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILN QIKR code 0000RGRQABG000000RG0000QGBR0000000R3PG00RG00000000QAAAAAAABG0RG000000RRG00000QAAAAAAAA B1BG LCS 0000 GRQABG000000RG0000Q BR0000000R 00RG00000 00QAAAAAAABG0RG000000RRG00000QAAAAAAAA B code 0000BGRQABG000000RG0000QABR0000000R00000RG00000R00QAAAAAAABG0RG000000RRG00000QAAAAAAAAABRRR 1npp ESVEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKRG site site + +++++++++ 1m1g GVK PSKVEFEKG DQVRVIEGPFMNFTGT VEEV HPEKRKLTVMISI FGRMTPVELDFDQVEKI code 0G0 00000000R 00000R00QB0R0000 R000 0RRRRG0R0O00R RR0R000000QABG0-- LCS 0 0 0 00R 00000 00QB0R0000 R000 0R G0 0 00 RR0 000000 0 code 0 0R0R1OG 00RG00000 00QB0R00000R00010R G0 0 003000RR03000000 0G-- 1npp V KPSKVEF EKGDQVRVI EGPFMNFTGTVEEVHPEK RK L TVMISIFGRMTPVELD FDQV site ++++++++++ *RATIO : lcs/trimmed_seq_a = 0.861 (= 205/ 238), lcs/trimmed_seq_b = 0.869 (= 205/ 236) |
| Structural alignment by ComSubstruct + manual inspection |
| DSSP .LLLLLEEEEEEELLLLHHHHHHHHHLHHHHHLLHHHEEEEELLLEEEEEEEELLLEEEEEELLLLLEEEEEELLLLEEEEEELLLLEEEE 1m1g .QELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVV |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1nnp vQELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVKVV DSSP lLLLLLEEEEEEELLLLHHHHHHHHHHHHHHLLLHHHEEEEELLLEEEEEEEELLEEEEEEELLLLLEEEEEELLLLEEEEEEELLEEEEE DSSP ELLLLLHHHLLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHLLLLEEEELEELLEELEELHHHHHHHHLLLLl 1m1g ESVEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKr ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1npp ESVEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIKr DSSP ELLLLLLHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHLLLLEEEELEELLEELEELHHHHHHHHHLLLl DSSP lllllllLLLLLLEEEELLLLLLLLEEEELLLLLlLLLEEEEEEELLEEEEEEELhHHEEel 1m1g gvkpskvEFEKGDQVRVIEGPFMNFTGTVEEVHPeKRKLTVMISIFGRMTPVELDfDQVEki ||||||||||||||||||||||||||| |||||||||||||||||||| 1npp gvkpskvEFEKGDQVRVIEGPFMNFTGTVEEVHPeKRKLTVMISIFGRMTPVELD.FDQV.. DSSP lllllllLLLLLLEEELLLLLLLLLEEEEEELLLlLLLEEEEEEELLEEEEEEEL.LLLL.. *Z-score 27.7 AR 230 RMSD 16.3A SI 98% |
| Structural alignment by DaliLite |
(1-6)
 |
 |
 |
Z-Score:
24.0
AR: 236 RMSD: 6.5A SI: 100% |
|
| 1m1g C (len 244) | 1m1g
D (len 239) |
DaliLite result | ||
| site 1m1C QVQELEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVK code --R1000000000000QGQAAAAAAAAAAAAABRRQABGR0000RG00000000RRG000000RG0000000000RR00000000ROG00 LCS 000000000Q QAAAAAAAAAAAAABR QABG 0000RG00000000R 000000RG0000000000R 00000000R G00 code --000000000QBQAAAAAAAAAAAAABRPQABG00000RG00000000R00000000RG0000000000R000000000RRG00 1m1D EKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVK site site 1m1C VVESVEGDTCVNAPPIS KPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQ I code 0R0001RGQAABG0000 01RG000OQABRG0000000RQG00RG00000R00QAAAAAAABG0RG000000RRG00000QAAAAAAAAH B LCS 0R000 R QAABG0000 0 RG000 QABR 0000000RQG00 G00000 00QAAAAAAABG0RG000000RRG00000QAAAAAAAA B code 0R0000R0QAABG0000G0 RG000RQABR00000000RQG000G00000000QAAAAAAABG0RG000000RRG00000QAAAAAAAAAAB 1m1D VVESVEGDTCVNAPPISKP GQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQIK site site +++ +++++++ 1m1C KRGVKPSKVEFEKGDQVRV IEGPFMNFTGTVEEVHPEKRKLTVMISIFG RMTPVELDFDQVEKI code GR0000R000000R00000 R00QB0R0000000000QBRG0000000RR G3000000QABG0-- LCS G 000 R00 00 R00000 R00QB0R0000000000QBR 00 0 00RR G 000000QABG0 code G 0001R00R00 R000000R00QB0R0000000000QBR 00R0R00RRRGR000000QABG0-- 1m1D R GVKPSKVEFE KGDQVRVIEGPFMNFTGTVEEVHPEKR KLTVMISIFGRMTPVELDFDQVEKI site ++++++++++ *RATIO : lcs/trimmed_seq_a = 0.883 (= 212/ 240), lcs/trimmed_seq_b = 0.902 (= 212/ 235) |
| Structural alignment by ComSubstruct + manual inspection |
| DSSP lllllLLEEEEEEELLLLHHHHHHHHHHHHHHHLLHHHEEEEELLLEEEEEEEELLLEEEEEELLLLLEEEEEELLLLEEEEEELLLLEE 1m1C qvqelEKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVK ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1D .....EKKWYALQVEPGKENEAKENLLKVLELEGLKDLVDEVIVPAEEKVVIRAQGKEKYRLSLKGNARDISVLGKKGVTTFRIENGEVK DSSP .....LLEEEEEEELLLLHHHHHHHHHHHHHHLLLHHHEEEEELLLEEEEEEEELLLEEEEEELLLLLLEEEEELLLLEEEEEELLLLEE DSSP EEELLLLLLHHHLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHLLLLEEEELEELLEELLLLHHHHHHHHHHH 1m1C VVESVEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQI |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1D VVESVEGDTCVNAPPISKPGQKITCKENKTEAKIVLDNKIFPGYILIKAHMNDKLLMAIEKTPHVFRPVMVGGKPVPLKEEEVQNILNQI DSSP EEELLLLLHHHLLLLLLLLLLEEEELLLLEEEEEEEELLLLLLEEEEEELLLHHHHHHHHHLLLEEEELEELLEELLLLHHHHHHHHHHH DSSP LLLLlllLLLLLLLLEEEELLLLLLLLEEEEEEEELLLLEEEEEEEELLEEEEEEEEHHHEEEL 1m1C KRGVkpsKVEFEKGDQVRVIEGPFMNFTGTVEEVHPEKRKLTVMISIFGRMTPVELDFDQVEKI |||| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||| 1m1D KRGVkpsKVEFEKGDQVRVIEGPFMNFTGTVEEVHPEKRKLTVMISIFGRMTPVELDFDQVEKI DSSP HHLLlllLLLLLLLLEEELLLLLLLLLEEEEEEEELLLLEEEEEEEELLEEEEEEEELLLLEEL *Z-score 24.0 AR 236 RMSD 6.5A SI 100% |
| Structural alignment by DaliLite |